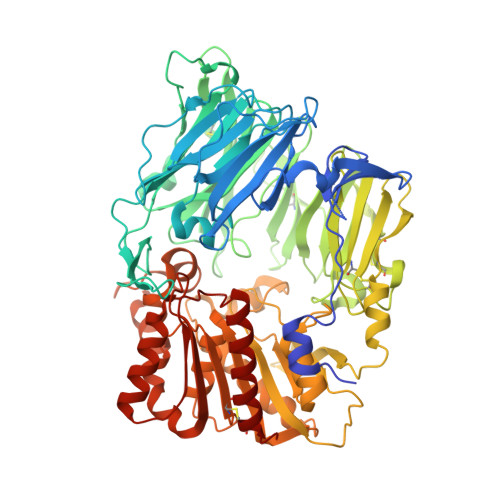
Structure-based design and synthesis of benzimidazole derivatives as dipeptidyl peptidase IV inhibitors.
Wallace, M.B., Feng, J., Zhang, Z., Skene, R.J., Shi, L., Caster, C.L., Kassel, D.B., Xu, R., Gwaltney, S.L.(2008) Bioorg Med Chem Lett 18: 2362-2367
- PubMed: 18346892
- DOI: https://doi.org/10.1016/j.bmcl.2008.02.071
- Primary Citation of Related Structures:
3CCB, 3CCC - PubMed Abstract:
A novel series of non-covalent, benzimidazole-based inhibitors of DPP-4 has been developed from a small fragment hit using structure-based drug design. A highly versatile synthetic route was created for the development of SAR, which led to the discovery of potent and selective inhibitors with excellent pharmaceutical properties.
Organizational Affiliation:
Takeda San Diego, Drug Discovery, 10410 Science Center Drive, San Diego, CA 92121, USA. michael.wallace@takedasd.com